A variety of genetic and genomic resources have been developed for the domestic cat. Each resources has added to the knowledge of the cat genome, aiding researchers in the development of the domestic cat as a model for human disease, improving the health of the feline itself, and also assisting forensic applications. All appropriate resources are welcome to be listed on this website – please submit addition resources that are novel or may have been overlooked.
Domestic Cat Karyotype
The domestic cat chromosomal complement is 2N = 38 (N = 19) with 18 autosomes and the XY sex chromosomes (XX is female, XY is male). Various cytogenetic techniques, such as R-, RBG-banding and fragile site studies, have also helped distinguish and characterize the cat chromosomes.47-49,52 For example, cats do not have a significant fragile X site on the X chromosome that is found in humans and is associated with mental retardation. Although a sequential numbering of the chromosomes has been suggested (Cho et al., 1997)3. This historical classification of chromosomes into morphologic groups has been retained in the cat.the classical chromosomal nomenclature that represents chromosomes by size and telomeric positions is still widely used to represent the cat karyotype and ideograms. Hence cats have three large metacentric chromosomes (A1 to A3), four large subtelomeric chromosomes (B1 to B4), two medium-size metacentrics (C1 to C2), four small subtelomerics (D1 to D4), three small metacentrics (E1 to E3), and two small acrocentrics (F1 and F2). The X chromosome is midsize and subtelomeric, similar to chromosome B4. All of Felidae have a similar karyotype to the domestic cat and the cat karyotype is highly representative of most carnivores.
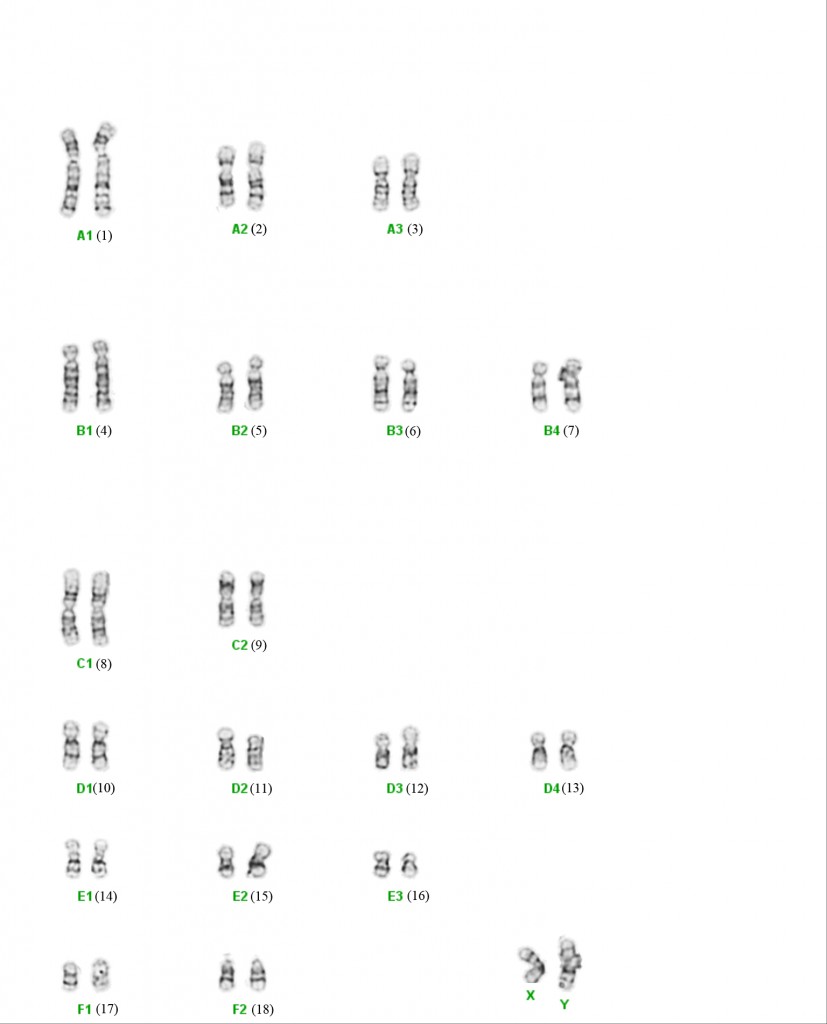
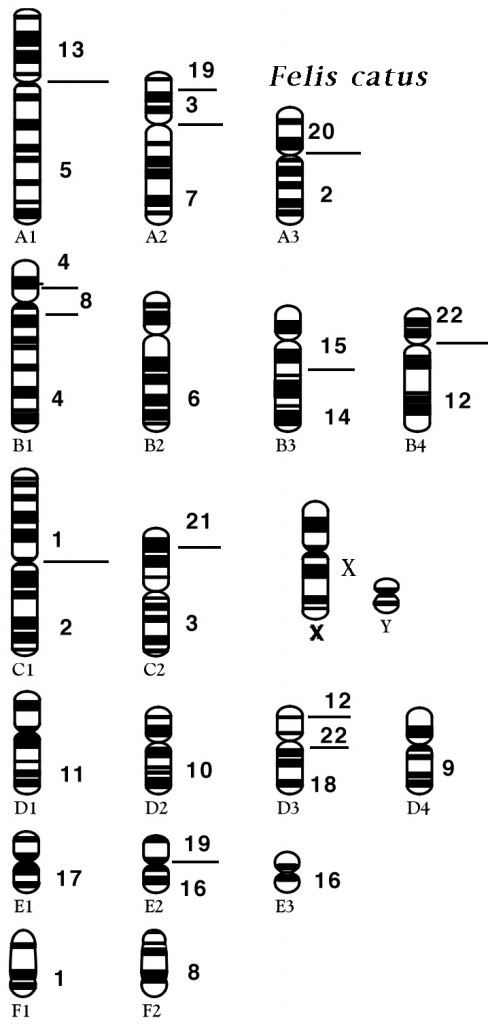
Early chromosome staining recognized some major alterations in the felid genome, particularly the Robertsonian translocation of F1 and F2 to form chromosome C3 in the ocelot lineage of cats from South America (2N = 36)59. Minor pericentric inversions, additions, or deletions of the small chromosomes cause variation in the felid karyotype; however, overall, domestic cats have a chromosomal architecture that is highly representative for all felids and ancestral for most carnivores. 34,44 The pericentric inversion of chromosome F1 produces a small, more centromeric chromosome and represents as E4 in many cat species.
Historically, the first genetic consideration to explain reduced fertility or intersex cats is chromosomal differences, especially the loss of one of the sex chromosomes. Karyotypic and now gene-based assays are common methods to determine if a cat with ambiguous genitalia50 or a poor reproductive history has a chromosomal abnormality. Karyotypic studies of male tortoiseshell cats have shown that they are often mosaics, or chimeras, being XX/XY in all or some tissues.* The minor chromosomal differences that are cytogenetically detectable between a domestic cat and an Asian leopard cat are likely the cause of fertility problems in the Bengal cat breed, which is a hybrid between these two species. Other significant chromosomal abnormalities causing common “syndromes” are not well documented in the cat.
*References 2, 4, 7, 10, 14, 19, 20, 42, 54.
Domestic Cat Chromosome Painting
The sufficient variation of cat chromosomal sizes also allowed for the easy flow sorting of cat chromosomes.56 Chromosome painting has indicated that the cat genome organization is highly conserved to that of human chromosomes. For example, the p arm of human chromosome 1(1p) is largely composed of the same genes that are on the cat chromosome defined as C1, whereas human chromosome 1q is composed of genes that are found on cat chromosome F1. The chromosome painting technique has also be performed reciprocally, implying painting cat chromosomes onto human mitotic chromosome spreads and human chromosomes onto cat mitotic chromosome spreads, revealing the high conservation of chromosomal arrangement of cat to humans,56,61 specifically compared to mice.53 Thus chromosome painting is an excellent overview of cat genome organization, 38 which greatly facilitates candidate gene approaches because the location of particular genes could be anticipated in cats from comparison with the genetic map of humans.55 This additional confirmation of conservation to human, with regard to genome organization, further supported additional genetic resource development for the cat as a valuable animal model for human disease.

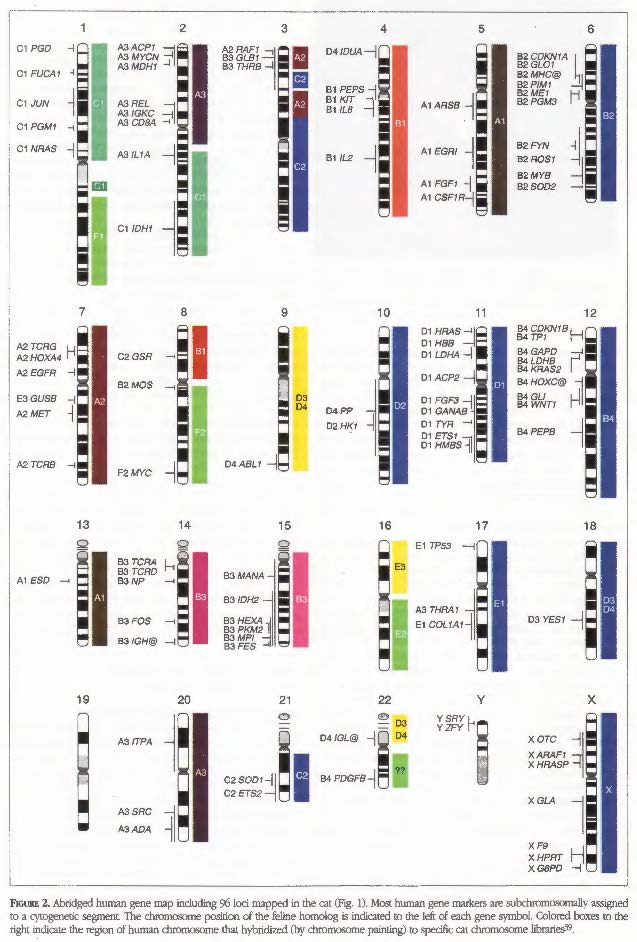
O’Brien SJ, Wienberg J, Lyon LA: Comparative genomics: lessons from cats,Trends Genet 13:393, 1997.
References
2. Centerwall WR, Benirschke K: Animal model for the XXY Klinefelter’s syndrome in man: tortoiseshell and calico male cats, Am J Vet Res 36:1275, 1975.
3. Cho KW, Youn HY, Watari T et al: A proposed nomenclature of the domestic cat karyotype,Cytogenet Cell Genet 79:71, 1997.
4. Chu EHY, Thuline HC, Norby DE: Triploid-diploid chimerism in a male tortoiseshell cat,Cytogenetics 24:1, 1964.
7. Doncaster L: On the inheritance of tortoiseshell and related colours in cats, Proc Camb Philol Soc13:35, 1904.
10. Gregson NM, Ishmael J: Diploid triploid chimerism in three tortoiseshell cats, Res Vet Sci 12:275, 1971.
14. Ishihara T: Cytological studies on tortoiseshell male cats, Cytologia 21:391, 1956.
19. Kosowska B, Januszewski A, Tokarska M et al: Cytogenetic and histologic studies of tortoiseshell cats, Med Weter 57:475, 2001.
20. Kuiper H, Hewicker-Trautwein M, Distl O: [Cytogenetic and histologic examination of four tortoiseshell cats], Dtsch Tierarztl Wochenschr 110:457, 2003.
34. Nash WG, O’Brien SJ: Conserved regions of homologous G-banded chromosomes between orders in mammalian evolution: carnivores and primates, Proc Natl Acad Sci U S A 79:6631, 1982.
38. O’Brien SJ, Wienberg J, Lyon LA: Comparative genomics: lessons from cats, Trends Genet 13:393, 1997.
42. Pyle RL, Patterson DF, Hare WC et al: XXY sex chromosome constitution in a Himalayan cat with tortoise-shell points, J Hered 62:220, 1971.
44. Rettenberger G, Klett C, Zechner U et al: ZOO-FISH analysis: cat and human karyotypes closely resemble the putative ancestral mammalian karyotype, Chromosome Res 3:479, 1995.
47. Ronne M: Localization of fragile sites in the karyotype of Felis catus, Hereditas 122:279, 1995.
48. Ronne M, Storm CO: The high resolution RBG-banded karyotype of Felis catus, In Vivo 6:517, 1992.
49. Ronne M, Storm CO: Localization of landmarks and bands in the karyotype of Felis catus,Cytobios 81:213, 1995.
52. Shibasaki Y, Flou S, Ronne M: The R-banded karyotype of Felis catus, Cytobios 51:35, 1987.
53. Stanyon R, Yang F, Cavagna P et al: Reciprocal chromosome painting shows that genomic rearrangement between rat and mouse proceeds ten times faster than between humans and cats,Cytogenet Cell Genet 84:150, 1999.
54. Thuline HC: Male tortoiseshell, chimerism and true hermaphroditism, J Cat Genet 4:2, 1964.
55. Wienberg J, Stanyon R: Chromosome painting in mammals as an approach to comparative genomics, Curr Opin Genet Dev 5:792, 1995.
56. Wienberg J, Stanyon R, Nash WG et al: Conservation of human vs. feline genome organization revealed by reciprocal chromosome painting, Cytogenet Cell Genet 77:211, 1997.
57. Wurster-Hill DH, Centerwall WR: The interrelationships of chromosome banding patterns in canids, mustelids, hyena, and felids, Cytogenet Cell Genet 34:178, 1982.
58. Wurster-Hill DH, Doi T, Izawa M et al: Banded chromosome study of the Iriomote cat, J Hered 78: 105, 1987.
59. Wurster-Hill DH, Gray CW: Giemsa banding patterns in the chromosomes of twelve species of cats (Felidae), Cytogenet Cell Genet 12:388, 1973.
60. Wurster-Hill DH, Gray CW: The interrelationships of chromosome banding patterns in procyonids, viverrids, and felids, Cytogenet Cell Genet 15:306, 1975.
61. Yang F, Graphodatsky AS, O’Brien PC et al: Reciprocal chromosome painting illuminates the history of genome evolution of the domestic cat, dog and human, Chromosome Res 8:393, 2000.