A variety of genetic and genomic resources have been developed for the domestic cat. Each resources has added to the knowledge of the cat genome, aiding researchers in the development of the domestic cat as a model for human disease, improving the health of the feline itself, and also assisting forensic applications. All appropriate resources are welcome to be listed on this website – please submit addition resources that are novel or may have been overlooked.
Domestic Cat Microsatellites (STRs)
The first cat microsatellite markers (short tandem repeats (STRs)) were developed in the Laboratory of Viral Carcinogenesis (later named Laboratory of Genomic Diversity – LGD) at the National Cancer Institute.34-35 These STRs were first localized by linkage using an interspecies backcross family (32) and later physically assigned and integrated into radiation hybrid maps (25,27,30-31,33).
These published markers have been developed into panels for parentage testing (21) and these and additional markers have been developed into panels for forensic applications (1,15,22) The same STRs have been used for several population studies of the cat as well (13, 17-18).
Additional novel STR markers (~500) that are closely linked to genes have been developed for the domestic cat but are yet unpublished and were based on assembly “felcat3”.
See this Excel file for Unpublished Cat gene-associated STRs.
STRs have been identified in silico from the genome sequence of the domestic cat (Version felcat4 (catChrV17e). To identify these STRS, di- and tetranucleotides repeats were selected as evenly spaced across all chromosomes, with repeat units 15 – 20 in length, and with a match score of 100%, implying a perfect repeat.
See this Excel file for Unpublished Cat dinucleotide in silico derived-STRs.
The distribution of the STRs can be visualized on this PDF.
See this Excel file for Unpublished Cat tetranucleotide in silico derived-STRs.
The distribution of the STRs can be visualized on this PDF.
Domestic Cat SNPs
Although a variety of mutations have been identified in the cat that are associated with traits and diseases, the first large-scale identification of SNPs occurred with the initial 1.9X genome sequencing of the cat, which identified SNPs within the one Abyssinian cat used in the sequencing effort (12), detecting ~327 k SNPs. A second Sanger-based effort for cat sequencing included the same Abyssinian cat and single representatives from several breeds, a domestic shorthair and an African wildcat (Felis lybica caffra)(2). This second effort increased the cat genome coverage to 2.8X total and identified ~964 K SNPs within the domestic cats and ~ 900 K SNPs between the domestic cats and the wildcat. A more in depth NextGen sequencing-based effort was implemented with NIH funding by the Genome Institute of Washington University at St. Louis – led by Wes Warren, PhD. Pools of 4 individual cats from six cat breeds (Birman, Egyptian Mau, Japanese Bobtail, Maine Coon, Norwegian Forest Cat, and Turkish Van) as well as a pool of random bred cats from Asia and a pool of wildcats, were included in the sequencing effort for SNP discovery. Over 17 million SNPs were discovered across these cat populations.
Domestic Cat DNA Array
In 2009, Hill’s Pet Food, Inc. donated $1,000,000 to the Morris Animal Foundation (MAF) for the development of genetic / genomic resources for the domestic cat. MAF developed a Cat Advisory Committee that suggested that the best use of these funds would be the development of a DNA SNP array for the domestic cat. The committee requested that Illumina, Inc. be commissioned to develop the cat array, which was released for use in February 2011. The donated funds produced 4,600 arrays that were awarded to the research community via competitive requests made to the Cat Health Network, which included support from the MAF, Winn Feline Foundation, American Veterinary Medical Foundation (AVMF) and the American Association fo Feline Practitioners (AAFP). GWAS and array-based studies are in progress for the domestic cat using the Illumina Infinium iSelect 63K Cat DNA Array.
The complete list of the SNPs on the Illumina Infinium iSelect 63K Cat DNA Array can be found in this Excel file “Cat array SNPs” and is available from a “Dropbox” that has been established by the Lyons Feline Genetics Laboratory at UC Davis. Request access to the “Dropbox” by sending an e-mail to Leslie A. Lyons, PhD – lyonsla@missouri.edu
- Cat DNA Array Design
To develop the cat DNA array, ~9.55 million SNPs from the three combined genome seqeuncing efforts were submitted to Illumina to produce an ~63K array. The submission criteria included: avoiding rare SNPs, SNPs near repeats or within duplication sites, SNPs with more than one allele and SNPs only found in Cinnamon, the Abyssinian cat used for the genome sequencing of the cat. Approximatley 5,000 SNPs that were identified in the wildcat were also submitted as well as phenotypic SNPs and SNPs that suggested phylogenetic importance in felids.
- Cat DNA Array SNP Quality
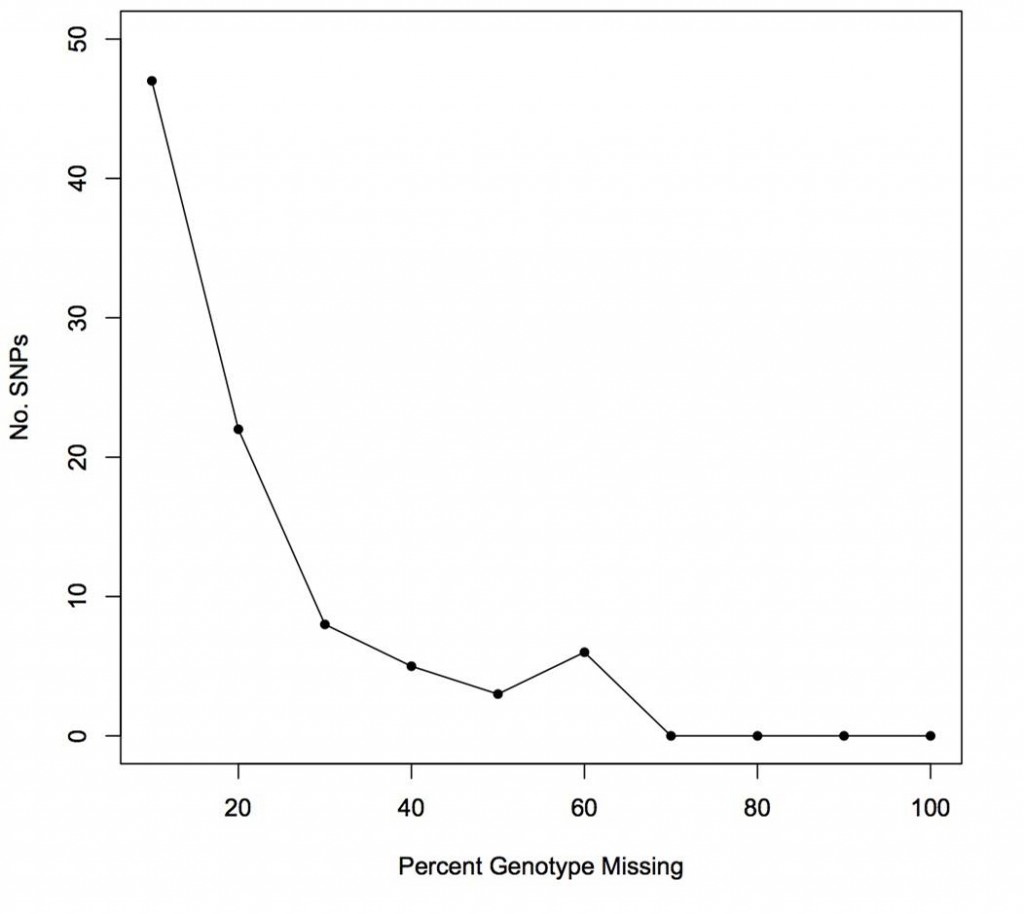
The Illumina Infinium iSelect 63K Cat DNA Array was tested using 288 cats from different 12 breeds, 10 wildcats, 10 western random bred cats and 10 eastern random bred cats. Five trios were tested, the Abyssinian (Cinnamon) and the 6 cats from the Hill’s SNP discovery project. The success of the ~63K SNPs on the array were highly successful, only a few hundred SNPs producing poor quality data.
| SNPs | No. | Fail | Note |
| Total | 62,897 | ||
| Domestic | 57,897 | ||
| Wildcat* | 4,200 | ||
| Autosomal | 53,140 | ||
| X-linked | 2,738 | ||
| Pseudoautosomal | |||
| Phenotypic | 35 | ||
| Phylogenetic | 92 (128) | ||
| Unassigned Contigs (38) | 6,891 | ||
| Popn. ID SNPS | 109 |
*Wildcat SNPs were distributed on different chromosomes, including the X and unassigned contigs.
| SNP Drop-out on Cat DNA Array per Breed | ||
| Breed* | SNPs MAF < 0.05 | Final Cats |
| Abyssinian | 22,080 | 18 |
| Birman (Sweden) | 22,317 | 20 |
| Burmese (USA) | 22,343 | 21 |
| Cornish Rex | 20,769 | 12 |
| Egyptian Mau | 20,281 | 15 |
| Japanese Bobtail | 16,330 | 13 |
| Maine Coon (Italy) | 16,836 | 20 |
| Norwegian Forest (Finland) | 12,795 | 18 |
| Persian | 19,119 | 21 |
| Ragdoll | 20,362 | 12 |
| Siamese | 23,518 | 19 |
| Turkish Van | 12,258 | 19 |
| Eastern Random | 11,007 | 19 |
| Western Random | 12,787 | 21 |
| Wildcat (F. silvestris) | 21,261 | 5 |
*Breed cats also included one Bengal and one American Curl within the 5 trios.
- Cat DNA Array Data (See Karotypes for Cat Chromosome No. Conversion)
GWAS and array-based studies are in progress for the domestic cat using the Illumina Infinium iSelect 63K Cat DNA Array. A “Dropbox” that has been established by the Lyons Feline Genetics Laboratory at UC Davis for access and sharing of the cat array data. Request access to the “Dropbox” by sending an e-mail to Leslie A. Lyons, PhD – lyonsla@missouri.edu. This“DropBox” includes an Excel file “Cat GWAS Signalment” that has the basic signalment of the cats used in the GWAS studies.
1: Menotti-Raymond M, David VA, Weir BS, O’Brien SJ. A population genetic database of cat breeds developed in coordination with a domestic cat STR multiplex. J Forensic Sci. 2012 May;57(3):596-601. doi: 10.1111/j.1556-4029.2011.02040.x. Epub 2012 Jan 23. PubMed PMID: 22268511.
2: Mullikin JC, Hansen NF, Shen L, Ebling H, Donahue WF, Tao W, Saranga DJ, Brand A, Rubenfield MJ, Young AC, Cruz P; NISC Comparative Sequencing Program, Driscoll C, David V, Al-Murrani SW, Locniskar MF, Abrahamsen MS, O’Brien SJ, Smith DR, Brockman JA. Light whole genome sequence for SNP discovery across domestic cat breeds. BMC Genomics. 2010 Jun 24;11:406. PubMed PMID: 20576142; PubMed Central PMCID: PMC2996934.
9: Schmidt-Küntzel A, Nelson G, David VA, Schäffer AA, Eizirik E, Roelke ME, Kehler JS, Hannah SS, O’Brien SJ, Menotti-Raymond M. A domestic cat X chromosome linkage map and the sex-linked orange locus: mapping of orange, multiple origins and epistasis over nonagouti. Genetics. 2009 Apr;181(4):1415-25. Epub 2009 Feb 2. PubMed PMID: 19189955; PubMed Central PMCID: PMC2666509.
10: Menotti-Raymond M, David VA, Schäffer AA, Tomlin JF, Eizirik E, Phillip C, Wells D, Pontius JU, Hannah SS, O’Brien SJ. An autosomal genetic linkage map of the domestic cat, Felis silvestris catus. Genomics. 2009 Apr;93(4):305-13. Epub 2008 Dec 13. PubMed PMID: 19059333; PubMed Central PMCID: PMC2656606.
12: Pontius JU, Mullikin JC, Smith DR; Agencourt Sequencing Team, Lindblad-Toh K, Gnerre S, Clamp M, Chang J, Stephens R, Neelam B, Volfovsky N, Schäffer AA, Agarwala R, Narfström K, Murphy WJ, Giger U, Roca AL, Antunes A, Menotti-Raymond M, Yuhki N, Pecon-Slattery J, Johnson WE, Bourque G, Tesler G; NISC Comparative Sequencing Program, O’Brien SJ. Initial sequence and comparative analysis of the cat genome. Genome Res. 2007 Nov;17(11):1675-89. PubMed PMID: 17975172; PubMed Central PMCID: PMC2045150.
13: Menotti-Raymond M, David VA, Pflueger SM, Lindblad-Toh K, Wade CM, O’Brien SJ, Johnson WE. Patterns of molecular genetic variation among cat breeds. Genomics. 2008 Jan;91(1):1-11. Epub 2007 Oct 25. PubMed PMID: 17964115.
15: Coomber N, David VA, O’Brien SJ, Menotti-Raymond M. Validation of a short tandem repeat multiplex typing system for genetic individualization of domestic cat samples. Croat Med J. 2007 Aug;48(4):547-55. PubMed PMID: 17696310; PubMed Central PMCID: PMC2080565.
16: Luo SJ, Johnson WE, David VA, Menotti-Raymond M, Stanyon R, Cai QX, Beck T, Yuhki N, Pecon-Slattery J, Smith JL, O’Brien SJ. Development of Y chromosome intraspecific polymorphic markers in the Felidae. J Hered. 2007;98(5):400-13. Epub 2007 Jul 23. PubMed PMID: 17646273.
17: Driscoll CA, Menotti-Raymond M, Roca AL, Hupe K, Johnson WE, Geffen E, Harley EH, Delibes M, Pontier D, Kitchener AC, Yamaguchi N, O’brien SJ, Macdonald DW. The Near Eastern origin of cat domestication. Science. 2007 Jul 27;317(5837):519-23. Epub 2007 Jun 28. PubMed PMID: 17600185.
18: Lipinski MJ, Froenicke L, Baysac KC, Billings NC, Leutenegger CM, Levy AM, Longeri M, Niini T, Ozpinar H, Slater MR, Pedersen NC, Lyons LA. The ascent of cat breeds: genetic evaluations of breeds and worldwide random-bred populations. Genomics. 2008 Jan;91(1):12-21. Epub 2007 Dec 3. PubMed PMID: 18060738; PubMed Central PMCID: PMC2267438.
19: Murphy WJ, Davis B, David VA, Agarwala R, Schäffer AA, Pearks Wilkerson AJ, Neelam B, O’Brien SJ, Menotti-Raymond M. A 1.5-Mb-resolution radiation hybrid map of the cat genome and comparative analysis with the canine and human genomes. Genomics. 2007 Feb;89(2):189-96. Epub 2006 Sep 25. PubMed PMID: 16997530.
21: Lipinski MJ, Amigues Y, Blasi M, Broad TE, Cherbonnel C, Cho GJ, Corley S, Daftari P, Delattre DR, Dileanis S, Flynn JM, Grattapaglia D, Guthrie A, Harper C, Karttunen PL, Kimura H, Lewis GM, Longeri M, Meriaux JC, Morita M, Morrin-O’donnell RC, Niini T, Pedersen NC, Perrotta G, Polli M, Rittler S, Schubbert R, Strillacci MG, Van Haeringen H, Van Haeringen W, Lyons LA. An international parentage and identification panel for the domestic cat (Felis catus). Anim Genet. 2007 Aug;38(4):371-7. PubMed PMID: 17655554; PubMed Central PMCID: PMC1974777.
22: Menotti-Raymond MA, David VA, Wachter LL, Butler JM, O’Brien SJ. An STR forensic typing system for genetic individualization of domestic cat (Felis catus) samples. J Forensic Sci. 2005 Sep;50(5):1061-70. PubMed PMID: 16225210.
25: Menotti-Raymond M, David VA, Agarwala R, Schäffer AA, Stephens R, O’Brien SJ, Murphy WJ. Radiation hybrid mapping of 304 novel microsatellites in the domestic cat genome. Cytogenet Genome Res. 2003;102(1-4):272-6. PubMed PMID: 14970716.
27: Menotti-Raymond M, David VA, Roelke ME, Chen ZQ, Menotti KA, Sun S, Schäffer AA, Tomlin JF, Agarwala R, O’Brien SJ, Murphy WJ. Second-generation integrated genetic linkage/radiation hybrid maps of the domestic cat (Felis catus). J Hered. 2003 Jan-Feb;94(1):95-106. Erratum in: J Hered. 2003 Nov-Dec;94(6):following table of contents. PubMed PMID: 12692169.
30: Sun S, Murphy WJ, Menotti-Raymond M, O’Brien SJ. Integration of the feline radiation hybrid and linkage maps. Mamm Genome. 2001 Jun;12(6):436-41. PubMed PMID: 11353390.
32: Menotti-Raymond M, David VA, Lyons LA, Schäffer AA, Tomlin JF, Hutton MK, O’Brien SJ. A genetic linkage map of microsatellites in the domestic cat (Felis catus). Genomics. 1999 Apr 1;57(1):9-23. PubMed PMID: 10191079.
33: Murphy WJ, Menotti-Raymond M, Lyons LA, Thompson MA, O’Brien SJ. Development of a feline whole genome radiation hybrid panel and comparative mapping of human chromosome 12 and 22 loci. Genomics. 1999 Apr 1;57(1):1-8. PubMed PMID: 10191078.
34: Menotti-Raymond M, David VA, Stephens JC, Lyons LA, O’Brien SJ. Genetic individualization of domestic cats using feline STR loci for forensic applications. J Forensic Sci. 1997 Nov;42(6):1039-51. PubMed PMID: 9397545.
35: Menotti-Raymond MA, David VA, O’Brien SJ. Pet cat hair implicates murder suspect. Nature. 1997 Apr 24;386(6627):774. PubMed PMID: 9126735.